Mutations
Do Mutations Drive Evolution?
In the evolutionary model, mutations are hailed as a dominant mechanism for pond-scum-to-people evolution and provide “proof ” that the Bible’s history about creation is wrong. But are we to trust the ideas of imperfect, fallible men about how we came into existence?
Mutations and New Genetic Information
If new genetic information—required to build eyes where there are none, for example—does not occur in nature, then evolution is stuck in the water. For evolutionists, the solution comes in the form of mutations. The problem is that the only beneficial mutations ever observed do not add new information to the genome.
Mutations in Viruses
This study illustrates the fact that natural selection can operate on a series of mutations and even on other organisms to produce a population change with a noticeably new characteristic. Yet for all that, the result was diversity within a kind.
Mutations Could Help in Finding HIV Vaccine
Human Immunodeficiency Virus (HIV) has frustrated efforts to create an effective vaccine. Analysis of naturally developed antibodies in a few infected patients, however, has uncovered a pattern in HIV mutations, a pattern that could be the key to developing an HIV vaccine.
Articles About Mutations
-
July 21, 2024 from Answers Magazine
When it comes to people with disabilities, Christians need to rethink what it means to be “fearfully and wonderfully made.”
-
Feb. 24, 2024 from Essays on Origins: Creation vs. Evolution
The survival of living species depends on its ability to pass on its genetic instructions, from generation to generation, without significant alteration.
-
July 22, 2022 from Answers in Depth
Did a bacteria colony just prove gain-of-function mutations exist?
-
Sept. 22, 2018 from Refuting Common Evolutionist Claims
Neo-Darwinism offers this basic equation for evolution: mutations + natural selection + millions of years = particles-to-person evolution.
-
Jan. 4, 2017 from Answers in Depth
If we share a common ancestor with a chimpanzee, as evolutionists confidently maintain, then how did our brains leap so far ahead in size and capability?
-
In-Depth ArticleJust How Random Are Mutations?Aug. 18, 2016 from Answers in Depth
Changes to the sequence of nucleotides (e.g., mutations) can alter the genetic information of the organism, which, in turn can alter its physical features
-
Book Chapter2.8 Mutation-Selection in Biblical PerspectiveMarch 28, 2016 from Creation: Facts of Life
Mutations are no real help in explaining the origin of species, but they are great for explaining the origin of disease, disease organisms, and birth defects.
-
Book Chapter2.5 Mutations, Yes; Evolution, NoMarch 28, 2016 from Creation: Facts of Life
Contrary to popular opinion, drug resistance in bacteria does not demonstrate evolution.
-
July 1, 2015 from Answers Magazine
Evolution would require an enormous amount of change. Modern laboratory experiments have tested bacteria’s ability to change. Is this ability truly unlimited?
-
In-Depth ArticleHijacking Good Science: Lenski’s Bacteria Support CreationAug. 13, 2014 from Answers in Depth
Lenski's long-term evolution experiment does not distinguish between observable limited change and unobservable molecules-to-man evolution.
-
De-Regulation of an Existing TraitOct. 6, 2012 from News to Know
Genome analysis confirms “a key evolutionary innovation” is only de-regulation of an existing trait.
-
Mutated Sense of TasteMarch 17, 2012 from News to Know
Many meat-eating mammals have mutated sweet sense.
-
Scientists Admit Genetic Data Timing UncertainMarch 17, 2012 from News to Know
“The inability to observe past mutation rates means that the timing of events from genetic data remains uncertain,” report Cambridge geneticists.
-
The Ancient Mega-CloneFeb. 11, 2012 from News to Know
Models suggest massive Mediterranean meadows are millenary mega-clones.
-
Mutational Secrets of a VirusFeb. 4, 2012 from News to Know
Series of mutations said to evolve a “key innovation”
-
Molecular Time-TravelJan. 14, 2012 from News to Know
“Molecular machine’s evolutionary trajectory” is an imaginary marvel.
-
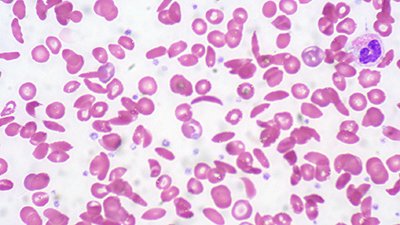
Link Between Sickle-Cell and Malaria ResolvedNov. 19, 2011 from News to Know
Mysterious malarial link to sickle-cell mutation resolved
-
Can Accumulated Mutations Produce New Species?Aug. 20, 2011 from News to Know
Can three walk together and evolve into one?
-
Genome Structure Over Genetic Mutations: What Makes You DifferentAug. 13, 2011 from News to Know
The genomic forest is harder to see than its trees.

Answers in Genesis is an apologetics ministry, dedicated to helping Christians defend their faith and proclaim the good news of Jesus Christ.
- Customer Service 800.778.3390
- Available Monday–Friday | 9 AM–5 PM ET
- © 2026 Answers in Genesis