Disease
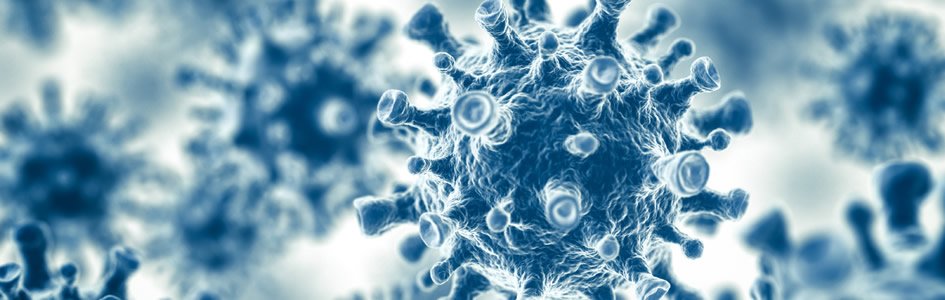
Swine Flu
Typically swine flu does not infect humans but occasionally it mutates and then is capable of infecting humans. Contact with pigs is usually necessary to “catch” swine flu but sometimes it mutates further and is transmissible from human to human.
Cancer
Cancer reminds us of the brokenness, the suffering, and the mortality of creation in this present age, all traceable back to Adam’s sin. Genesis makes it clear that everything in the original creation, was “very good.” We can infer that cancer was not a part of that, since the Bible describes death as an “enemy.”
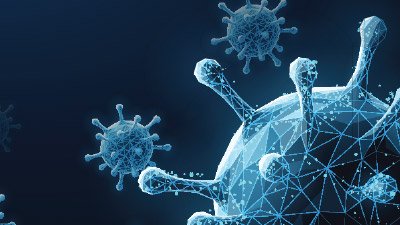
Bird Flu
It is plain to see that the genetic code of bird flu viruses does not stay the same but is constantly mutating and rearranging. Because virus proteins interact with proteins in the cells they infect, changes in the viral protein can have effects on what type of cells can be infected. So what should one say if asked, “Is the ‘bird flu’ evolving?”
What About Malaria and Other Diseases Before the Fall?
Mosquitoes in the original creation were likely pollinators able to derive the nutrients needed to reproduce from plants without consuming blood. But what about the malaria-causing parasite Plasmodium before “the Fall”?
Disease Topics
Articles About Disease
-
Aug. 20, 2025 from Answers in Depth
Mosquitoes may be annoying pests, but they are also the deadliest creatures in our sin-cursed world.
-
Feb. 28, 2024 from Answers Magazine
When our oldest son was diagnosed with Williams syndrome—a rare genetic disorder that causes cognitive and physical disabilities—my husband and I found ourselves in a whole new world.
-
Aug. 25, 2023 from Answers in Depth
Did God create parasites to make us sick?
-
Aug. 4, 2021 from Answers in Depth
Tick-borne diseases like Lyme disease have plagued humans for millennia. Did God make them that way, and what do they teach us about a fallen creation?
-
July 3, 2021 from Answers in Depth
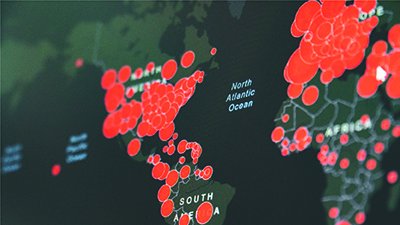
A creation perspective on why God allows emerging and reemerging diseases like COVID-19 and Lyme Disease that cause emotional, financial, and health problems.
-
Nov. 22, 2017 from Answers in Depth
Understanding the flu’s origins will practically aid us in treating the flu virus, so let’s take a look first at what causes the flu.
-
In-Depth ArticleThe Design of the Mosquito and Its DangersAug. 11, 2016 from Answers in Depth
If everything that God made was good, where did disease-causing mosquitoes come from? What is the origin of mosquito-borne diseases?
-
July 1, 2016 from Answers Magazine
Zika, spread by mosquitoes, can severely damage unborn babies’ nervous systems and cause them to be born with tiny heads (microcephaly).
-
Evolutionary Conjecture Cannot Cure CancerNov. 27, 2015 from Countering the Culture
Evolutionary scientists assume the patterns they see result from evolution and therefore erroneously believe evolutionary thinking can help the war on cancer.
-
Nov. 22, 2015 from Answers Magazine
Every second, our bodies must monitor a melange of threats that seek to invade and destroy.
-
In-Depth ArticleSuper Staph in the Community: Is It Evolving?Oct. 22, 2015 from Answers in Depth
Is Community-associated (CA)-MRSA “evolution in action”? Microbiology research based on creation provides some answers to its emerging dominance in the USA.
-
Book ChapterIs Antibiotic Resistance Proof of Evolution?July 3, 2015 from The Genesis of Germs
Antibiotic resistance is one of the most important topics that a beginning biology student going into medicine should learn and understand.
-
May 14, 2015 from Answers in Depth
Fleas are considered a nuisance. How can they be explained as a part of God’s very good creation?
-
Jan. 24, 2015 from Answers in Depth
Is the war on malaria plagued by “rapid evolution of insecticide resistance” in mosquitoes?
-
Nov. 1, 2014 from Answers in Depth
A physicist and an astrobiologist team up to explain to medical doctors how knowledge of evolution holds the key to curing cancer.
-
Technical In-Depth ArticleThe Genesis of MalariaJune 19, 2013 from Answers in Depth
Microbiology and parasitology research based on the creation paradigm appears to provide some answers puzzling questions.
-
Parasites Affect Behavior of MothsSept. 17, 2011 from News to Know
Gypsy moth virus packs a weapon of mass dispersion.
-
Grandpa CancerApril 2, 2011 from News to Know
Cancer reminds us of brokenness, suffering, and mortality traceable to Adam’s sin. But evolutionists propose that cancer may be our ancestor!
-
Oct. 25, 2009 from Answers Magazine
Leprosy has terrified humanity since ancient times and was reported as early as 600 BC in India, China, and Egypt. Hansen’s disease is still a major health problem.
-
Magazine Department ArticleFlu AwayOct. 1, 2009 from Answers Magazine
H1N1 flu virus is an example of mutation, not evolution in the usual sense of the word.

Answers in Genesis is an apologetics ministry, dedicated to helping Christians defend their faith and proclaim the good news of Jesus Christ.
- Customer Service 800.778.3390
- Available Monday–Friday | 9 AM–5 PM ET
- © 2026 Answers in Genesis