Chapter 2
Bacteria Structure
Section 2
The basic elements of bacterial anatomy include (1) the cell wall and (2) the inner cell body, or cytoplasm. The latter is surrounded by a cytoplasmic membrane just inside the cell wall, and commonly contains various granules, other cell inclusions, and vitally important nuclear material. In addition, the entire bacterium may be enclosed in an envelope or sheath of slimy, viscous or gelatinous material, forming a more or less definite capsule. There may be organs of locomotion called flagella, and some bacilli have pili. Certain bacteria may develop internally special structures called spores.
The basic elements of bacterial anatomy include (1) the cell wall and (2) the inner cell body, or cytoplasm. The latter is surrounded by a cytoplasmic membrane just inside the cell wall, and commonly contains various granules, other cell inclusions, and vitally important nuclear material. In addition, the entire bacterium may be enclosed in an envelope or sheath of slimy, viscous or gelatinous material, forming a more or less definite capsule. There may be organs of locomotion called flagella, and some bacilli have pili. Certain bacteria may develop internally special structures called spores.
Cell Walls and Gram Stain
Since the majority of bacteria are not spherical, instead maintaining distinct shapes, such as that of a long rod, it is obvious that there must be an outer wall of considerable rigidity. Such a wall cannot ordinarily be seen in preparations of bacteria colored with commonly used stains, but at times the cytoplasm of a bacterium becomes contracted, pulling away from the wall, revealing the latter clearly. The existence of this cell wall, distinct from both the outer slime layer (capsule) and the cytoplasmic membrane, has been clearly demonstrated by special staining and electron micrographs. From the behavior of living bacterial cells in motion, it is inferred that the cell wall is rather rigid, with little elasticity. The Gram stain was named for Danish physician Hans Christian Gram, who discovered a technique that could differentiate all known bacteria into two categories. The staining technique, utilizing various dyes, would leave Gram-positive cells a deep purple and Gram-negative cells bright red. During the late 1800s and early 1900s, the reason for this distinction was unclear. By the mid-1900s, electron microscopy of bacteria revealed differences in the fine structure of cell walls of Gram-negative and Gram-positive bacteria. Gram-negative walls are not continuously associated with the underlying cytoplasmic membrane, have a thick layer of peptidoglycan, and usually are multi-layered. In Gram-positive bacteria, the walls are usually closely associated with the cytoplasmic membrane and can consist of a single, amorphous layer or of several layers—the structure varying in different species.
Through the impetus of investigating the mechanism of antibiotic action, great advances have been made in understanding bacterial cell-wall chemistry. Both Gram-positive and Gram-negative cell walls contain the same chemical substance forming the backbone of the rigid structure. This material, peptidoglycan, is a complex polymer containing two amino sugars, glucosamine and muramic acid (Latin murmus, “wall”) and several amino acids. The muramic acid in bacterial cell walls is unique; no other type of cell is known to contain this sugar.
These simple components join to form this complex and interwoven cross-linked chemical structure. Peptidoglycan is responsible for the rigidity of bacterial cell walls. Some Gram-positive organisms contain teichoic (Greek teichos, “wall”) acids important for pathogen specificity. Other bacteria have special attached carbohydrates or proteins, such as the M-protein of streptococci, contributing to its virulence. Gram-negative bacteria contain lipopolysaccharides in their cell walls that sometimes contain an endotoxin. The cell walls of Gram-negative bacteria are more complex chemically than those of Gram-positive organisms. They contain lipoproteins, lipopolysaccharides, and phospholipids, but contain no teichoic acid. Only four or five different amino acids are found in Gram-positive cell walls. Their lipid content is minimal, but they can contain teichoic acid.
Cell Membrane and Membrane Systems
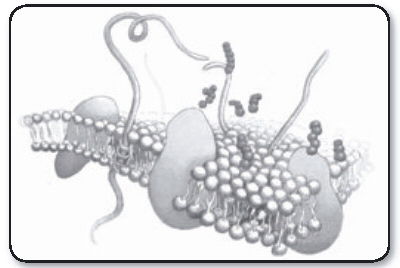
A bacteria cell membrane. In some bacteria that cause strep throat (i.e. Streptococcus pyogenes), their cell membrane and envelope components have M-proteins that make the throat feel scratchy and sore.
The plasma membrane (or cell membrane) of eukaryotic and prokaryotic cells is similar in function and basic structure. There are, however, differences in the protein types of the membranes. Eukaryotic membranes contain carbohydrates that serve as receptor sites, assuming roles in such functions as cell-to-cell recognition. These carbohydrates also provide attachment sites for bacteria. Plasma membranes carry enzymes for metabolic reactions such as nutrient breakdown, energy production, and synthesis.
Bacteria contain a network of interconnected, membranous structures in their cytoplasm. Mesosomes are invaginations of the cytoplasmic membrane into the cytoplasm (right). They are irregular infoldings of plasma membrane artifacts, but not true cell structures. These structures serve not only as sites for respiratory enzymes, but also as regulators of orderly division of bacteria. Mesosomes provide a continuous membrane linking together nuclear fibrils at one end with the cytoplasmic membrane at the other. Since division involves both of the latter structures, mesosomes may be the means for their coordination during division. Ribosomes of bacteria are thought not to exist in the cytoplasm as discrete particles, but arranged in linear fashion on interconnected membranes attached to the plasma membrane.
The Nuclear Region
The nuclear apparatus in bacterial cells is obscured when cells are stained by basic dyes because of the presence in the cytoplasm of relatively large amounts of RNA, which has affinity for these dyes. These structures are sometimes called chromatin bodies, but in reality are nuclei and are also seen through the electron microscope. They have been shown to divide just before the whole bacterial cell divides. Chemically, the nuclei contain DNA bound to a basic protein. Bacterial “nuclei” are not surrounded by a membrane, as in true eukaryotic nuclei. The nuclear substance consists of DNA arranged into bundles of fibers. These bundles can rapidly change from compact arrangements into long, tape-like forms, depending on the environment.
Capsules, Glycocalyx, and Biofilms
A distinct layer often surrounds many bacterial cells beyond the cell wall. In some cases it is referred to as a capsule, and at other times it is a less defined, adhesive layer called a glycocalyx. This layer often releases a sticky, gelatinous, enveloping slime layer. The slime layer serves as an adhesive and growth medium for more bacteria to reproduce. This enveloping sheath is too thin to be seen in most species, but in some it regularly develops into a clearly visible capsule external to the cell wall, with a sharply defined outer edge following the contour of the cell body. Often the capsule is much wider than the bacterial cell itself. Chained or paired organisms are often enclosed within a single, continuous capsule.
The capsular material may arise from a modification of the cell wall, or may be secreted by the living bacteria, remaining attached as a firm, mucilaginous structure. Some bacteria secrete glycocalyx into their surroundings without forming a definite capsule. Capsule formation is markedly influenced by environmental conditions; it can be induced, in special circumstances, in strains of bacteria ordinarily showing no evidence of any slime layer or biofilm. These biofilms can form not only on natural environmental structures like rocks in a stream, but also contact lenses, food bowls, used syringes, catheters, urinary-tract tissue, and human skin and tongues. Biofilm microbes are estimated to be 1,000 times more resistant to microbiocides. Disease experts estimate that 65 percent of human bacterial infections involve biofilms.
The bacterium that causes anthrax, Bacillus anthracis, has generated much interest recently. The capsule of the anthrax bacillus is a polypeptide, consisting mainly of D-glutamic acid. Among pathogenic bacteria, the phenomenon of capsule formation is of prime importance. Species that also develop large capsules include Streptococcus pneumoniae, Klebsiella pneumoniae, and Bacillus anthracis. Most pathogenic bacteria develop at least a small amount of capsular substance when growing in body tissues, and the most conspicuous capsules are seen on organisms freshly obtained from the infected host. The capsules seem to act as a defense against bactericidal factors in body fluids. Therefore, they contribute directly to the disease-producing power, or virulence, of the organisms. These capsules are lost by most species after cultivation for some time in the laboratory, usually resulting in markedly lost capacity to produce disease.
Encapsulation is also significant in another respect. The chemical structure of the capsular material determines the specificity of protective antibodies formed by the infected individual. The numerous “types” of pneumococci, for example, differ from one another in the chemical composition of the polysaccharides making up their capsules. Consequently, each type stimulates the formation of a different antibody. Immunity to a particular type of pneumococcus requires an antibody that will react with the capsulated organisms of that type.
Motility and Flagella
All known spirilla, and about half the more familiar species of bacilli, possess the power of locomotion through liquids; that is, they can move independently from place to place. Bacteria possessing this capacity are said to be motile; those that cannot move about independently are non-motile. The movement of motile bacteria can be plainly seen when examined under the microscope. For their size, these mighty microbes are some of the fastest swimmers in the Designer’s creation. At top speed, an E. coli can swim nearly 50 times the length of its body in one second inside your urinary tract; the equivalent of a six-foot man swimming at 200 mph. Yet E. coli devotes less than two percent of its energy to swimming from your bladder toward the kidney inside the ureters. It achieves this remarkable feat through a special propulsion system, a speedboat-like “motor” turning spiral propellers at up to 100,000 revolutions per minute. The reversible motors are controlled by a complex feedback system that monitors the concentrations of food and/or toxins outside the bacteria, adjusting the motors accordingly. Bacteria do not run on gas or electricity the way man-made outboard motors do; the Master Engineer must have constructed these incredible microbes in motion. (See Design Focus 2.2 on flagella function.)
The organs of locomotion are delicate, hair-like processes extending from one or more parts of the cell bodies of the motile bacteria. These are called flagella (Latin for “little whips”). They do not have a lashing movement, however. Instead, the organisms appear to be propelled by wavelike, rhythmical contractions passing along the length of the flagella (10 to 20 μ). The flagella filament arises from the cytoplasm, anchored by a basal region consisting of a hook-like structure and a basal body. The basal body has a central rod and set of enclosing “O rings.” They are usually somewhat coiled and often much longer than the bacterium from which they arise. They are remarkably thin, having a width of only about 0.013μ, and are easily broken off. Apparently, when the bacteria are in motion, the flagella often entwine to form a tail-like locomotor organ. The pattern of flagellar attachment to bacterial cells, as well as the number of flagella, is distinct to each species. In Pseudomonas, a single flagellum, or a tuft of flagella, is located at the end of the bacterium as polar flagella; E. coli has multiple flagella projecting from all around the cell known as peritrichous flagella.
Chemical analyses indicate that flagella are composed mostly of fibrous protein subunits termed flagellin. This protein is contractile and similar to the contractile proteins in muscle tissue. There are nearly 50 proteins making up all the subunits associated with flagella, and most of these are highly interdependent. The subunits of flagellin aggregate into long strands that twist about each other in a helical fashion to form the flagella. Two, three, or as many as five strands, depending on the species of bacteria, can form the helices. Flagellin apparently undergoes elastic contractions and expansions to account for flagellar movement. Bacterial flagella are unique structures not equivalent to the cilia or flagella of protozoa or higher organisms.
Focus on Flagella Function
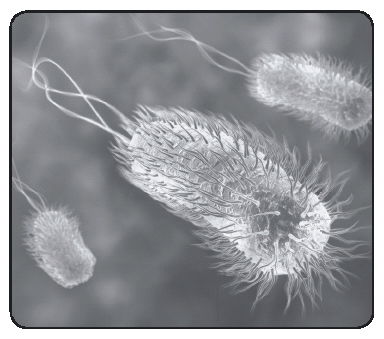
Computer artwork of E. coli bacteria showing flagella
Antony van Leeuwenhoek, using a single-lens microscope, was intrigued by “animalcules” (little animals) that he saw in his well water. He wanted to know whether they might survive exposure to pepper, so he ground up some and added it to a sample. The number of animalcules waxed and waned until August 6, 1676, when he made a discovery: “I now saw very plainly that these were little eels, or worms, lying all huddled up together and wriggling; just as if you saw, with the naked eye, a whole tubful of very little eels and water, with the eels a-squirming among one another: and the whole water seemed to be alive with these multifarious animalcules. This was for me, among all the marvels that I have discovered in nature, the most marvellous of all; and I must say, for my part, that no more pleasant sight has ever yet come before my eye than these many thousands of living creatures, seen all alive in a little drop of water, moving among one another, each several creature having its own proper motion.”
When Leeuwenhoek described little or “minute eels,” he was looking at a spirillum, probably Spirillum volutans, the large bacterium shown in the accompanying sketch. Leeuwenhoek never saw its flagella. Flagella, the organelles of locomotion were first seen on Chromatium okenii, another large bacterium, by Christian Ehrenberg in 1836, and later on S. volutans by Ferdinand Cohn in 1872. The role that flagella play in that response was examined in detail after dark-field condensers were developed, beginning in 1909 with work done by Karl Reichert and culminating in 1920 with the work of Paul Metzner, who described the motion of flagellar bundles of S. volutans in stunning detail.
S. volutans has two flagellar bundles, each composed of about 25 flagellar filaments. Here, the cell is swimming from left to right. Its body is helical. The bundle on the left is in the tail configuration; the one on the right is in the head configuration. When the filaments change their directions of rotation, the bundles switch their configurations and the cell moves in the opposite direction. An Escherichia coli bacterium is shown below the S. volutans for comparison. As many as six flagellar filaments arise at random from the sides of the E. coli cell and form a bundle that appears near one pole. Rotation of the filaments in the bundle pushes the cell forward. When the bundle changes its orientation, the cell goes off in a new direction.
Bacteria may have one flagellum or many, displayed on the surface in a variety of patterns. Polar flagella come off the ends of bacteria, whereas peritrichous flagella are distributed randomly over the entire surface (peri means “around”; trichous means “hair”). Bacteria with polar flagella may have one, two, or even a tuft of hundreds of flagella at one or both ends of the cell. A few bacteria called spirochetes have internalized flagella that lie beneath the cell wall and coil around the cytoplasmic membrane.
The way bacterial flagella move is completely different from the action of eukaryotic flagella. Bacterial flagella neither flex nor whip; they rotate like the propellers on a boat. If a bacterium with a single polar flagellum were held by its flagellum so that the flagellum did not move, then the body of the whole bacterium would rotate. Rotation is accomplished by a basal body and the hook that connects the flagellum to the bacterial cell. The basal body attaches the base of the flagellum to the cytoplasmic membrane and cell wall and acts as a motor to turn the flagellum. The hook transfers the rotation from the basal body to the external flagellar filament.
Flagella rotate counterclockwise to propel bacteria forward, and the motion is driven by chemotaxis, which is the movement of bacteria in response to chemicals in the environment. Especially important are chemicals that can be used as energy sources, and bacteria have receptors on their surface to detect these molecules. When such a molecule interacts with the receptor, a signal is sent to the basal body, the flagellar motor starts, the flagellum rotates, and the bacterium moves toward the energy source.
Bacteria with a single polar flagellum move simply back and forth. They move forward by rotating their flagellum counterclockwise, and in reverse by rotating their flagellum clockwise. How do bacteria with a tuft of polar flagella or flagella distributed peritrichously over their entire surface move? Won’t the flagella get all tangled up? In order for bacteria like these to move, the action of all the flagella must be synchronized.
Bacteria with peritrichous flagella or tufts of polar flagella show an overall pattern of movement consisting of a series of “runs” (or “swims”) and “tumbles” (or “twiddles”). During a run, all of the flagella are brought together into a functional bundle and rotated synchronously in a counterclockwise direction to propel the bacterium toward the energy source. During tumbles, the flagellar bundles disassemble. The relative proportion of time spent in runs determines how fast the bacterium moves in a specific direction and depends on the concentration of the energy source. The greater the concentration of molecules of energy source, the more interaction there is with the receptors, the more time the flagella rotate counterclockwise and the farther the bacterium moves. As the concentration of the energy source decreases, there is less interaction with the receptors, the flagella are engaged less, and the bacterium does not move as far. Instead, the bacterium tumbles more often. During a tumble, flagella turn clockwise, but rather than reversing the direction of the movement, this causes bacteria to cease forward motion and randomly jiggle about. Every time a bacterium finishes a tumble, it moves randomly away from the site where the tumble occurred. Thus, the more a bacterium tumbles, the greater the chance that it will not move in a definite direction. Motility requires that more time be spent in runs than in tumbles.
Pili and Fimbriae
A number of species of Gram-negative bacilli possess surface, hair-like structures distinct from flagella. These are sometimes called pili or fimbriae. A single hair or a few long hairs are usually referred to as pili, while multiple, short, hair-like structures are called fimbriae. Pili have no relation to motility, occurring in both non-motile and motile organisms. They are very thin fibrils shorter than flagella, numbering several hundred on a single bacterial cell. They can be visualized only in electron micrographs. Pili may serve as a means for obtaining nutrients and attaching bacilli to surfaces. In liquid media cultures, pili may aid in forming a surface skin, or pellicle, of growth. Pili apparently are composed of subunits of a protein called pilin. These subunits probably aggregate into fibers that in turn wind about each other to form a hollow, helical, tube-like structure.
Design Focus 2.2
E. coli — “Superbug” and Evidence of Extraordinary Engineering
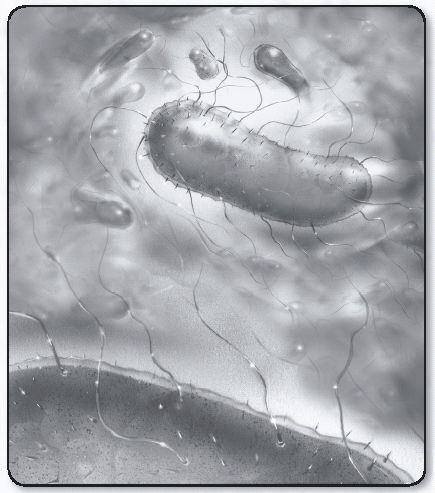
Illustration of E. Coli
Just how simple are bacteria? For decades, biology students have learned the unscientific concept that prokaryotic cells are “primitive.” Creation scientists counter that if it is alive, it is necessarily complex. Only those with a Darwinian worldview maintain that microorganisms such as bacteria are simple or basic. For more than half a century, Escherichia coli has been the tireless workhouse of biological research. In the 1940s, it was used as a host organism to determine the life cycle of viruses. Many of the important metabolic pathways, including the renowned Krebs cycle, were first worked out in this organism. In the 1950s, biochemists used E. coli to discover the three forms of microbial recombination. In the 1960s, it was the major research organism for deciphering the genetic code and learning how genes work. In the 1970s, E. coli became the guardian of public health as a valuable indicator of water pollution. It also emerged as an industrial giant, producing enzymes, growth factors, and vitamins. Since the 1980s, biochemists have used it as a living factory to produce an array of genetically engineered pharmaceuticals. In the 1990s, E. coli continued to illustrate how bacteria can be put to work in the interest of science and for the betterment of humanity.
E. coli is a testament to complexity. After its genome was completed, a technical review of the microbiology literature on its genetic and DNA design inferred that its systems were irreducibly complex in nature. E. coli has intricate regulation and transport systems, and other complex cellular processes. It is an important bacterial species normally found in the large intestine (unless one is taking antibiotics). It is one of the most thoroughly, intensely studied of all microorganisms and is found anywhere there are people.
Then along came E. coli 0157:H7. To be sure, this strain has caused much human misery because of its propensity to invade intestinal tissues, pass to the blood, and cause serious kidney damage. It has made us more careful about what we eat, causing us to think twice about having a rare hamburger. Along the way it has become the “germ of the week” on various news programs, portrayed as the chief villain in a world of villains. Unfortunately, this has made us forget all the good things E. coli has done for us. What a shame! Perhaps one bad apple can indeed spoil the whole barrel. Unless we are very hygienic, non-lethal strains can be found even on our hands.
Evolutionists traditionally have viewed bacteria as simple forms of life. These single-celled organisms, so the story goes, were one of the first life forms to have naturalistically sprung from the primeval oceans several billion years ago, and therefore have to be of simple construction. However, contrary to evolutionary assumptions, research has shown bacteria to be highly organized. We have also witnessed the discovery of new phenomena, such as autoaggregation of chemotactic bacteria and coordinated behaviors in complex-colony morphogenesis.
Bacterial cells are highly ordered, with communication and decision-making capabilities enabling them to coordinate growth, movement, and biochemical activity. Are these not amazing processes for a “simple” organism?
Just how “primitive” is E. coli? Biologists still are not sure. Even five years after the E. coli genome was announced, microbiologists are still far from knowing all the details of how the cell operates, lives, replicates, coordinates, and adapts to changing circumstances. There are tens of thousands of journal articles on E. coli, and new biological information continues to accumulate. New metabolic capabilities are discovered, connected to underlying genes. There are new regulation systems, new transport systems, and more information on cellular constituents and processes, but how many regulators are needed to maintain coordination of expression of the genes and proper interaction among gene products? Regulation systems are not the same in all bacteria, and we still do not have all the information for the regulatory networks of even one bacterial species. The minimal set of genes and proteins necessary to sustain an independently replicating cell does not have an easy answer.
Experimentation into details of the biology of E. coli continues to grow, yet not all enzymes and pathways in E. coli are known. Besides data for genes for unknown enzymes, we have data for enzymes that don’t have genes. There have been 55 enzymes of E. coli isolated, purified, and characterized over the years, but many of their genes have never been identified. The advent of DNA-sequencing technology and the completion of more than 50 microbial genomes now available to the public have not yet brought us to a complete understanding of exactly how a single, free-living cell functions and adapts to changing environments. Creation scientists cheerfully predict that if and when science does have a complete understanding of how a free-living cell functions and adapts, evolutionism will have nothing to do with it!
Conclusion
Autoaggregation? Regulatory networks? Transport systems? Coordinated behavior? Communication and decision-making capabilities? Enhanced export abilities? Is this why we should believe bacteria evolved from non-living chemicals? One could just as easily be speaking of a massive, high-tech automated factory — a sophisticated organization that certainly doesn’t spring up by chance, time, and natural processes! This is another example of how evolutionary theory contradicts science.
Why do evolutionists insist on preaching that bacteria (prokaryotes) are simple or primitive, when empirical research shows the opposite? Worse, how can they say that “simple” life evolved from non-life, using nothing but chance and time? As one creation biologist has said, when people make things smaller and smaller, it’s considered a wonder of modern technology (nanotechnology). When “nature” does it, it’s primitive. The Christian microbiologist can look through a creation lens in his microscope and observe an awesome, distinct design in E. coli!
The Genesis of Germs
Professor Alan Gillen shows that constantly mutating diseases are proof for devolution rather than evolution.
Read OnlineRecommended Resources

Answers in Genesis is an apologetics ministry, dedicated to helping Christians defend their faith and proclaim the good news of Jesus Christ.
- Customer Service 800.778.3390
- Available Monday–Friday | 9 AM–5 PM ET
- © 2026 Answers in Genesis